The activities are as follows:
- Teacher Guide
- Student activity, Graph Type A, Level 3
- Student activity, Graph Type B, Level 3
- Student activity, Graph Type C, Level 3
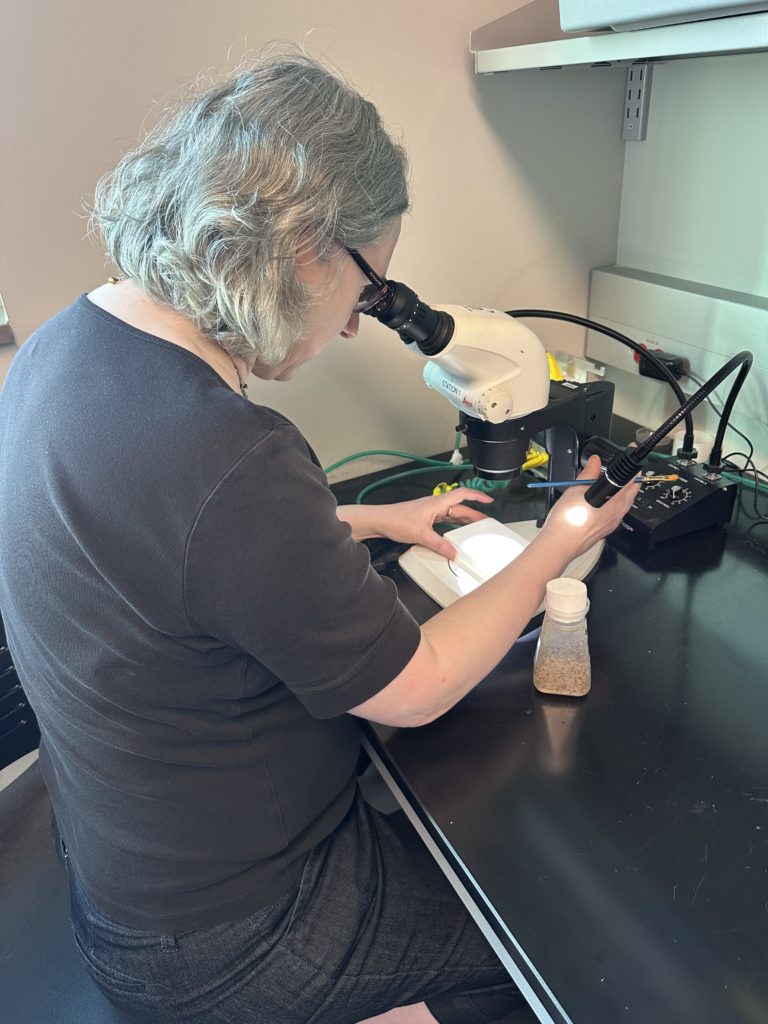
- Nicole Riddle Scientist Profile
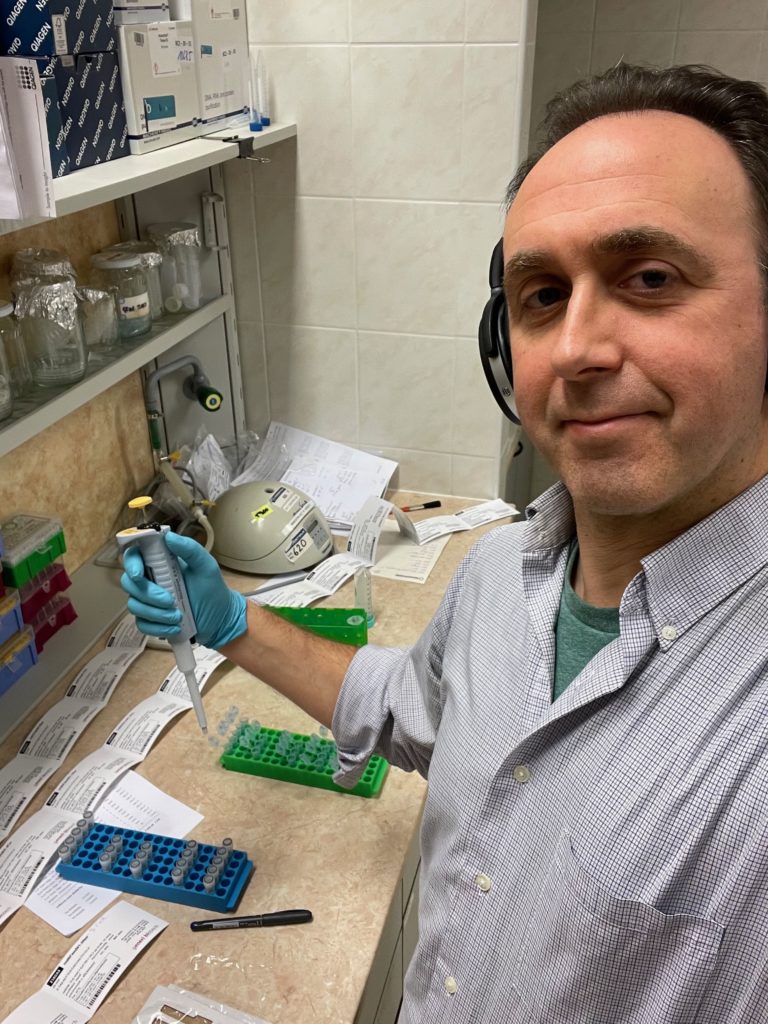
- Jamie Walters Scientist Profile
- Grading Rubric
Many factors affect lifespan, or how long an organism lives. Different species, and individuals within a species, will all live to different ages. Across species, things like body size, rate of metabolism, and genetics can all come into play. For example, larger animals tend to live longer than smaller organisms. Within a species, genetics and environmental conditions, such as being able to find food, the presence of predators, and disease, will also impact survival.
Scientists have also noticed that in many animal species, one sex tends to live longer than the other. Sometimes it is the males, and sometimes it is the females. Why might this be? To better understand aging differences across sexes, a group of scientists decided to work together. Each scientist studies a different species, so by combining their knowledge, they can look for patterns and see if there are consistent factors that are the cause.
Nicole and Jamie are two scientists in this group. Nicole studies fruit flies, while Jamie studies moths and butterflies. Even though fruit flies and moths are both insects, sex is determined differently. In most animals, biological sex is determined by specific chromosomes. These structures are inside cells and carry genetic information. Individuals usually have two sex chromosomes. Whether those two chromosomes are the same or different often determines whether their bodies develop as male or female.
In fruit flies, females have two of the same sex chromosomes (XX), while males have two different sex chromosomes (XY). In moths and butterflies, the pattern is reversed. Males have two of the same sex chromosomes (ZZ), while females have two different ones (ZW).
Nicole and Jamie wondered if having two different sex chromosomes might affect lifespan. When an individual has only one copy of a particular chromosome—like the X in XY males or the Z in ZW females—there is no second copy for the genes on that chromosome. If that single copy contains a harmful mutation or becomes damaged, the organism cannot rely on a second copy to make up for it. On the other hand, individuals with two of the same sex chromosomes (XX or ZZ) have a kind of “genetic backup”. This extra protection might reduce the risk of problems that could lead to an earlier death.
To test their idea about sex chromosomes and lifespan, Nicole and Jamie designed an experiment called a survival assay. A survival assay is a laboratory experiment in which scientists carefully track how long organisms live under controlled conditions. By keeping the environment consistent, scientists can focus on the specific factor they want to study.

Plodia interpunctella female by Pekka Malinen, Luomus is licensed under CC BY-SA 4.0.
Nicole performed her survival assay with the fruit fly species, Drosophila melanogaster. Jamie worked with a pantry moth species called Plodia interpunctella. Both scientists already raise these species in their labs and carefully document the life cycles and age of each individual.
To set up their assays, Nicole and Jamie chose individuals that had emerged from the pupae stage around the same time. This step was important because they wanted to make sure all individuals had the same starting point. If some individuals had emerged a lot sooner, the results would not be accurate.
Nicole collected 100 female and 100 male fruit flies, and Jamie collected 60 male and 60 female moths. The insects were given plenty of food and kept in good environmental conditions, such as appropriate temperature and humidity. By reducing stress, they could better observe natural lifespan differences between males and females, rather than differences caused by harsh conditions.
Each day, Nicole and Jamie recorded how many males and females were still alive. This careful daily tracking allowed them to see how survival changed over time. The survival assay continued until the last individual had died. By the end of the experiment, Nicole and Jamie had detailed data showing how long males and females lived in each species. These results would help them test whether having two identical sex chromosomes—or two different ones—might influence lifespan.
Featured scientists: Nicole Riddle (she/her) from the University of Alabama at Birmingham and Jamie Walters (he/him) from the University of Kansas.
Flesch–Kincaid Reading Grade Level = 9.9
Additional Teacher Resources:
- Scientist profiles: Nicole Riddle and Jamie Walters both have scientist profiles to supplement this activity. Have students read more about their research, personal lives, and advice they have as a way to share contemporary scientist role models with students!
- You can learn more about the IISAGE (Integration Initiative: Sex, Aging, Genomics, and Evolution) project here. This initiative is a collaborative effort to learn more about the mechanisms of sex-specific differences in aging and features research with a variety of organisms.
- Visit this page for additional scientist profiles and Data Nuggets featuring IISAGE research.